Spatial and temporal control of mRNA localization and translation plays critical roles in many events during germ cell development and asymmetric body pattern formation. Aberrant mRNA localization and untimely protein synthesis during oogenesis can lead to defect and mis-patterning of germ cell formation, causing embryonic lethality and sterility. One well-studied example is the assembly of germ plasm in Drosophila embryogenesis.
Oskar is a critical component in the regulation of Drosophila pole plasm formation and assembly. Like many other mRNAs, osk mRNA is not translated until properly localized in the oocyte. The posterior localization signal of osk mRNA is situated in the 3’-UTR region. Translation of osk mRNA is repressed by RNA interference at an early stage and later by the binding of protein factors to the 3’-UTR, and is alleviated after localization by the binding of activators. Once Osk protein is synthesized and accumulated in the posterior pole, it recruits other proteins and RNAs, such as Vasa Tudor, Aubergine and nanos, to co-localize in the posterior area. The place where osk mRNA is localized and translated will develop into germ plasm.
Although the function and the mechanism of oskar mRNA localization have been studied in great detail, the role of Oskar protein in germ-line formation and body patterning remains poorly understood. To explore the detailed mechanism of how Oskar protein interacts with other components to exert its function, the team of researchers lead by Dr. XU Rui-Ming at Institute of Biophysics (IBP) of Chinese Academy of Sciences (CAS) have determined the 3D structures of an N-terminal LOTUS/OST-HTH domain (Osk-N) and a C-terminal hydrolase-like domain of Oskar (Osk-C).
They firstly identified Oskar as a novel RNA-binding protein. They found that Osk-C has a lipase-fold structure but lacks critical catalytic residues at the putative active site. Surprisingly, they discovered that Osk-C has RNA binding function and can bind to the 3’UTRs of oskar and nanos mRNAs. This is the first time that a lipase-like domain has been reported to bind nucleic acids. Three crucial arginine residues forming a narrow positively charged ridge in the surface of Osk-C, and they are shown to be important for RNA binding. The researchers also identified Osk-N as a homodimer of winged-helix-fold modules, but without detectable RNA-binding activity.
These findings provided structural insights into how a hydrolase-like RNA binding domain may interact with RNA and suggest possible functions of Oskar in the regulation of translation and localization of relevant mRNAs through direct interaction with their 3’UTRs. These discoveries are supposed to direct future mechanistic studies of Oskar for a better understanding of its function in germ cell development.
This work was published online on August 31st 2015 by Proceedings of the National Academy of Sciences of USA. The work was a result of collaboration between Dr. XU’s group at IBP, CAS and Dr. Ruth Lehmann at New York University School of Medicine.
The research was supported by grants from the Ministry of Science and Technology of China, the National Natural Science Foundation of China and CAS. Dr. YANG Na, who is the first and co-corresponding author, is also supported by the Youth Innovation Promotion Association of the Chinese Academy of Sciences. Dr. YU Zhenyu and Mr. HU Menglong are the co-first authors in this publication.
Full Text
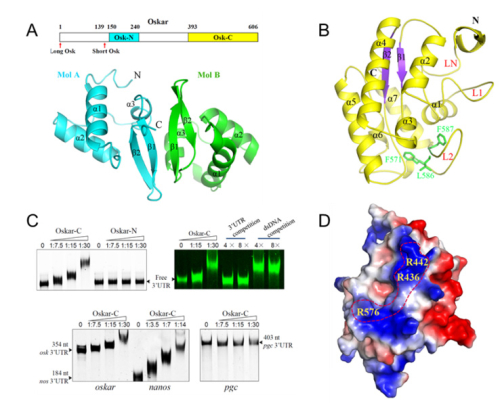
Fig. The structures of Osk-N and Osk-C and their RNA binding properties. (A). A schematic drawing of domain organization of Osk and overall structure of Osk-N homodimer; (B). Overall structure of Osk-C; (C) EMSA analyses of Osk-RNA interactions; (D). Important residues of Osk-C form a narrow positively charged ridge, which may be involved in RNA binding. (image by XU Rui-Ming's group at IBP)
Contact:
XU Rui-ming
Institute of Biophysics, Chinese Academy of Sciences
Website: http://english.ibp.cas.cn/ibp_pi/XY/201312/t20131230_115002.html
Email: rmxu@ibp.ac.cn
YANG Na
Institute of Biophysics, Chinese Academy of Sciences
Website: http://english.ibp.cas.cn/pe/casyipam/201406/t20140617_122898.html
Email: yangna@moon.ibp.ac.cn