R-M systems are almost universal and exist in ~90% bacterial and archaeal genomes. Type I restriction enzymes were the first REases to be purified and have been studied for five decades, which has helped launch the molecular biology revolution. Atomic structures of Type I R-M enzyme complexes have been difficult to obtain. Till now, only the crystal structures of individual subunits have been solved, but not of complexes. Despite half a century of efforts, the molecular details of the interactions among different subunits and the dynamic conformational transitions are still unclear due to the lack of high-resolution structures.
Professor YAN Xiao-Xue cooperation with Professor GAO Pu in the Institute of Biophysics of the Chinese Academy of Sciences determined the crystal structure of the Type I MTase complex from Thermoanaerobacter tengcongensis bound to a non-target DNA in the "open" conformation. The intermolecular interactions between the S and M subunits are mediated by a newly identified four-helix bundle formed by the CRs of the S subunit and the C-terminal alpha helices of the two M subunits. The four-helix bundle, rather than the TRD domains, determines the specificity of the interaction between the S and M subunits. The linker region between the C-terminal alpha helix and the N-terminal global domain of the M subunit is essential to control the conformational transition of the MTase complex. Further, biochemical results show that the R subunits prefer to load onto the "closed" form MTase rather than the "open" form.
To our knowledge, this is the first report of the atomic structure of the Type I R-M enzyme complex. we proposed an updated model for the complex assembly and conformational transition of the Type I R-M system. The structural and biochemical characterization of the Type I R-M system reported in this study provides guidelines for future applications in molecular biology.
This work is entitled “Structural basis underlying complex assembly and conformational transition of the Type I R-M system ”and published online by the Proc Natl Acad Sci USA on Oct 2, 2017. This research was supported by grants from the Natural Science Foundation of China and the Strategic Priority Research Program of the Chinese Academy of Sciences. The X-ray diffraction data were collected at the BL-17U/ BL-19U1 beamline at SSRF.
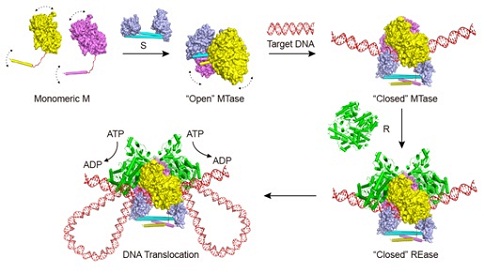
Figure: The proposed model for the complex assemblyand conformational changes of the type I R-M systems.
Contact
Yan Xiao-Xue
National Laboratory of Biomacromolecules
Institute of Biophysics, Chinese Academy of Sciences
Tel:86-10- 64888509(office)
E-mail: snow@ ibp.ac.cn
Or
Gao Pu
CAS Key Laboratory of Infection and Immunity
Institute of Biophysics, Chinese Academy of Sciences
Tel:86-10- 64887199(office)
E-mail: gaopu@ ibp.ac.cn