In a study published in Nature, Dr. Li Guohong and Dr. ZHU Mingzhao from the Institute of Biophysics of the Chinese Academy of Sciences and the collaborators demonstrated that the histone variant H2A.Z facilitates licensing and activation of early replication origins.
DNA replication is a tightly regulated process that ensures the precise duplication of the genome during cell proliferation. Replication origins determine where replication starts at the genome and regulate the whole genome replication program. There are tens of thousands of origins in the human genome; however, only about 10% of them are used in each cell cycle. How are the origins selected? In eukaryotes, DNA wraps octamers to form chromatin in the nucleus. The licensing and activation of replication origins are regulated by both DNA sequence and chromatin features. However, the chromatin-based regulatory mechanisms remain largely uncharacterized.
The scientists first found that knocking down H2AFZ genes in HeLa cells results in cell growth defects. Through mass spectrometry, they found that many subunits of prereplication complex were enriched on H2A.Z mono-nucleosomes, which indicating that H2A.Z may be involved in the licensing of DNA replication origins.
To find the mechanism of the interaction between H2A.Z and prereplication complex, scientists performed biochemistry analysis in tubes, and found that H2A.Z-containing nucleosomes bind directly to the histone lysine methyltransferase enzyme SUV420H1, promoting H4K20me2 deposition, which further recruit ORC1 (Origins recognition Complex 1), to help to accomplish the licensing of DNA replication origins.
Next, scientists confirmed the role of H2A.Z on DNA replication through genome-wide studies in HeLa cells. They found that signals of H4K20me2, ORC1 and nascent DNA strands (indicating active DNA replication origins) co-localize with H2A.Z, and the depletion of H2A.Z results in decreased H4K20me2, ORC1 and nascent-strand signals. In addition, H2A.Z-regulated replication origins have a higher firing efficiency and early replication timing compared with other origins.
To study the function of H2A.Z-regulated replication in a more physiological context, scientists conditionally knocked out (CKO) H2az1/H2az2 in T cells by generating CD4CreH2A.Zf/f mice. They found that in H2A.Z CKO mice, the activated T cells have defects of cell proliferation and DNA replication.
Thus, this study provides a novel epigenetic regulation mechanism of DNA replication origin selection and offers a new orientation on DNA replication regulation in eukaryote. In addition, this regulatory pathway can potentially be targeted for treatment of cancer and regulation of T cell function during immunotherapy.
This work has published in Nature on Dec 25th, 2019 titled “H2A.Z facilitates licensing and activation of early replication origins”. 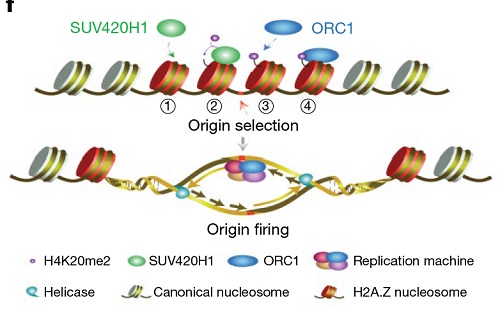
Origin selection: H2A.Z nucleosomes bind Suv420H1 directly (①) to establish H4K20me2 on chromatin (②), which then recruits ORC1 (③) to bind to replication origins (④); Origin firing: H2A.Z-Suv420H1-H4K20me2-ORC1 axis selectively license and activate early replication origins. Working model (The image by Dr. LI Guohong’s lab)
The web link for this paper is https://www.nature.com/articles/s41586-019-1877-9 Contact: LI Guohong
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
Phone: 86-10-64856269
Email: liguohong@ibp.ac.cn (Reported by Dr. LI Guohong's group) |