Chromatin compacts DNA to an extreme extend and allows eukaryotic genome fit the size of the nucleus. On the other hand, however, it must process the ability to untighten DNA and to permit the cellular machinery access to genome. Chromatin consists of nucleosomes in which a protein core is constituted by four canonical histones H2A, H2B, H3, H4 and wrapped around by 147 bp of DNA. Histone variants, and the chromatin remodelling machinery, can reorganize the compaction of chromatin and thus be important for epigenetic regulation of gene expression.
Histone variant H2A.Z is a universal mark of dynamic nucleosomes. H2A.Z is essential for growth, development and viability of a number of species including mammals. H2A.Z plays critical roles in multiple biological processes including gene transcription and replication, DNA repair, and genome integrity. The chromatin incorporation of H2A.Z is catalysed by SRCAP, an ATP-dependent, multi-component chromatin remodelling complex. The YL1 subunit of SRCAP is essential for securing H2A.Z recognition.
A new study published online by Nature Structural and Molecular Biology on March. 14, revealed a mechanism by which the histone variant H2A.Z is preferentially recognized by SRCAP chromatin-remodelling subunit YL1. A combination of biochemical, biophysical and yeast genetic analyses reveals the mechanism by which YL1 protein specifically interacts with histone variant H2A.Z rather than canonical histone H2A. The study elucidated a novel mechanism for histone variant recognition that is essential for H2A.Z replacement pathway.
LIANG Xiaoping and his colleagues from the research group led by Prof. ZHOU Zheng at Institute of Biophysics, Chinese Academy of Sciences solve the structure of H2A.Z-H2B dimer in complex with YL1. The 1.9-Å resolution complex structure reveals that an YL1-Z domain locating at the N-terminal YL1 clasps over H2A.Z–H2B. Three specific residues in H2A.Z confer the preference for YL1-Z and are essential for H2A.Z functions.
This study has been done in collaboration with Prof. Carl Wu at HHMI Janelia Institute. It was supported by grants from the Ministry of Science and Technology of China and the National Natural Science Foundation of China.
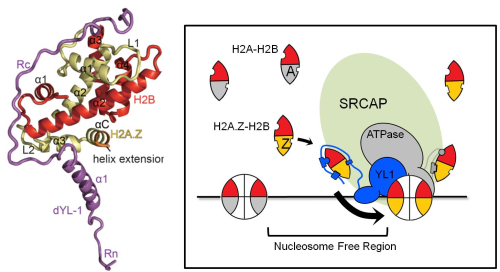
Figure: The structure of YL1-Z in complex of H2A.Z-H2B (left) and the graphic model of H2A.Z recognition and transfer mediated by SRCAP (right). (Iamge by IBP)
Full Text
CONTACT:
ZHOU Zheng
Laboratory of Infection and Immunity, Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
Phone: 010-64889862
Email: zhouzh@ibp.ac.cn