Aging, the major risk factor for a wide range of chronic diseases, is determined by both genetic and epigenetic factors. Accurate technology to track the chromatin dynamics in live cells is required for understanding the deeper mechanisms of aging process. So far, several technologies, including fluorescent in situ hybridization (FISH), TALE and CRISPR/dCas9 system, have been utilized to image specific loci in human genome. However, they are limited by non-live cell imaging, low signal to noise ratio and transfection efficiencies, as well as the formation of non-specific protein aggregates in the transfected cells.
In a new study as a cover story published in Cell Research on April 7, 2017, entitled “Visualization of Aging-Associated Chromatin Alterations with an Engineered TALE System”, researchers from Institute of Biophysics, Chinese Academy of Sciences (CAS) and their partners developed a new 3D genome imaging technique named as TTALE (by fusing thioredoxin to TALE), with which precisely visualizing specific genomic repetitive-sequence loci became possible.
In the current study, researchers discovered that conventional TALEs are prone to form insoluble aggregates, hampering their utility for precise genomic imaging in all human cell lines examined. To overcome this barrier and by screening a panel of peptides/proteins that are known to facilitate the expression of insoluble proteins in E. coli and fusing them to TALE, researchers found that the fusion of the thioredoxin to TALE (designated as TTALE) effectively eliminated the aggregate-like signals from conventional TALE. TTALEs were tested in a variety of cellular contexts, including human cancer cell lines, human embryonic stem cells, iPSCs and differentiated cells, as well as mouse cells in vitro or in vivo. “TTALEs can be applied to various human cell types and mouse cells, with imaging quality comparable to 3D-FISH and transfection efficiency better than the CRISPR/dCas9 system,” says Guang-hui Liu, a professor in Institute of Biophysics, CAS and senior author of the paper.
Importantly, for the first time, the location of 28S rDNA inside nucleolus in live cells was precisely visualized by TTALE system, which may provide a powerful tool for exploring the function of ribosomal DNA and nucleolus in different biological processes. In addition, MUC4 as a protein-coding gene locus can also be precisely labeled by TTALE in human cells. “Their success with this latter locus (MUC4) emphasizes the potential utility of the TTALE system for a wide range of experimental questions.” says Brian Kennedy, the President and CEO of the Buck Institute for Research on Aging in Novato, California.
Chromatin disorganization has been linked to human aging and age-related diseases. Although telomere shortening has been known as a hallmark of aging, repeat sequences known as the dark matter in human genome were rarely studied for their function to regulate biological processes including aging. This study identified rDNA attrition as a hallmark of aging in three models of human mesenchymal stem cell aging, as well as in the peripheral blood cells isolated from aged human individuals. “These initial findings open the possibility that the rDNA repeats may be a common source of genome instability and a driver of aging across diverse eukaryotes”, says Brian Kennedy.
Although the finds in this study focused on the ability of TTALEs to explore the chromatin dynamics with age, the application of TTALEs could be extended far beyond, such as studying the single nucleotide polymorphisms-induced pathologies like autism-related disorders and detecting the chromosomal or cell cycle aberrations in cancer. As Brian K. Kennedy says, “TTALEs promise to provide a robust platform for imaging chromatin dynamics under physiological conditions, as well as a potential tool for genome editing in therapeutic approaches to various pathologies.”
This work was supported by National Basic Research Program of China (973 Program), the Strategic Priority Research Program of the Chinese Academy of Sciences, Beijing Natural Science Foundation, National High Technology Research and Development Program of China, National Natural Science Foundation of China, Program of Beijing Municipal Science and Technology Commission, Key Research Program of the Chinese Academy of Sciences.
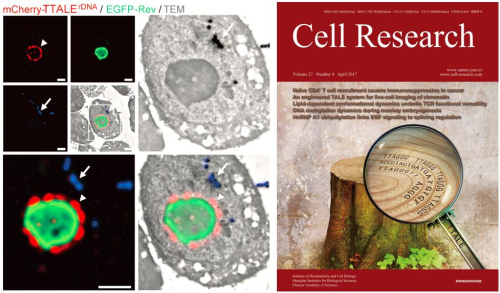
Figure1. TTALE-based precise imaging of 28S rDNAs in nucleolar organizer region (left), and the cover of Cell Research (right). (Image by IBP)
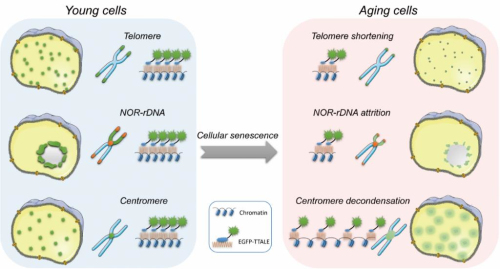
Figure2. Visualization of aging-associated alterations in genomic repetitive elements by TTALE systems indicating physical attrition of telomeric DNA repeats and NOR-rDNA repeats, as well as decondensation of centromeric DNAs. (Image by IBP)
Contact:
LIU Guang-hui; XU Tao
National Laboratory of Biomacromolecules
Institute of Biophysics, Chinese Academy of Sciences
86-10-64889970
QU Jing
State Key Laboratory of Stem Cell and Reproductive Biology, Institute of Zoology, Chinese Academy of Sciences
Email: ghliu@ibp.ac.cn; qujing@ioz.ac.cn; xutao@ibp.ac.cn