Stepwise tRNA Recognition Drives T-Box Riboswitch Regulation
T-box riboswitches are an important class of gene-regulatory elements whose function critically depends on their three-dimensional structure and conformational dynamics. However, the dynamic mechanism by which T-box riboswitches accurately recognize tRNAs with different aminoacylation states and subsequently initiate transcriptional or translational regulation has remained unclear.
Recently, a research team from the Institute of Biophysics of the Chinese Academy of Sciences, and Tsinghua University applied single-molecule fluorescence resonance energy transfer (smFRET) to systematically investigate the structural dynamics of the Mycobacterium tuberculosis ileS T-box riboswitch.
This study uncovered the molecular mechanism by which the riboswitch discriminates the aminoacylation status of the tRNA 3′ end during co-transcription, thereby regulating the initiation of translation.
This work was published in Nature Communications on 25 November 2025.
By selecting multiple labeling pairs within the T-box aptamer domain and discriminator domain, and combining UBP-based site-specific fluorescent labeling with smFRET analysis, the researchers comprehensively monitored the structural transitions of the T-box riboswitch induced by binding of tRNAs in different aminoacylation states.
They found that the T-box achieves two-step recognition-identity verification and aminoacylation-state sensing-through a conformational selection mechanism.
To further explore how co-transcriptional processes influence T-box regulatory mechanisms, the researchers generated T-box truncations of various lengths (simulating distinct transcriptional intermediates) as well as fluorescently labeled co-transcriptional T-box/tRNA complexes, and performed additional smFRET measurements.
The results showed that recognition of the tRNA NCCA terminus by the T-box is strongly dependent on transcriptional progression. This co-transcriptional regulation enables the riboswitch to rapidly respond to intracellular amino acid availability, ensuring efficient and precise control of gene expression.
Based on these findings, the researchers proposed a co-transcriptional regulatory model for the T-box riboswitch, elucidating the complete dynamic process through which it sequentially recognizes tRNA, senses aminoacylation, undergoes conformational switching, and ultimately modulates gene expression during transcription.
This study provides important mechanistic insights into RNA-RNA interactions and offers a theoretical foundation for the development of new antibiotic strategies.
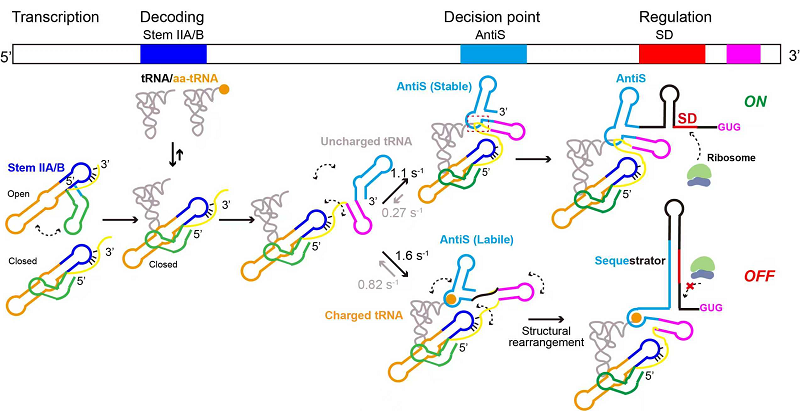
Figure: Model of Co-transcriptional Gene Regulation by the T-box Riboswitch
(Image by FANG Xianyang's group)
Article link: https://doi.org/10.1038/s41467-025-65388-6
Contact: FANG Xianyang
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
E-mail: fangxy@ibp.ac.cn
(Reported by Prof. FANG Xianyang's group)

