Scientists Map Spatiotemporal DNA Methylation Dynamics During Mammalian Post-Implantation Development
A research team led by Prof. LIU Jiang at the Institute of Biophysics, Chinese Academy of Sciences, reports SmC-seq, a spatial DNA methylation sequencing approach that enables genome-wide 5mC profiling at single-cell-scale resolution while preserving tissue context. The method is applied to generate a spatiotemporal atlas of DNA methylation during mammalian post-implantation development, and is published in Nature Methods on April 27, 2026.
Obtaining genome-wide DNA methylation profiles at single-cell resolution while preserving spatial context in intact tissues has remained technically challenging. SmC-seq addresses this by integrating microfluidic barcoding with enzymatic 5mC conversion chemistry. The method further incorporates an in situ histone removal step, reducing chromatin-associated constraints on enzymatic reactions and enabling an uniform genome-wide coverage.
SmC-seq performs in situ methylome profiling on tissue sections at ~10 μm resolution, corresponding to single-cell scale. In terms of throughput, the method captures on the order of 10,000 spatially indexed pixels per experiment. While preserving spatial information, SmC-seq achieves data quality comparable to that of existing single-cell DNA methylation sequencing approaches lacking spatial resolution.
The method is applied to investigate mammalian post-implantation development. The data reveal a layered methylation architecture in the placenta, consistent with its functional organization. Notably, genome-wide DNA demethylation is observed in maternal decidual cells following embryo implantation. These cells are proposed to differentiate into a lineage that provides nutrient support for mammalian yolk sac formation, offering insights into the origins of yolk sac development.
As a generalizable platform for spatial epigenomic profiling, SmC-seq expands the toolkit for studying development, aging, and disease in situ.
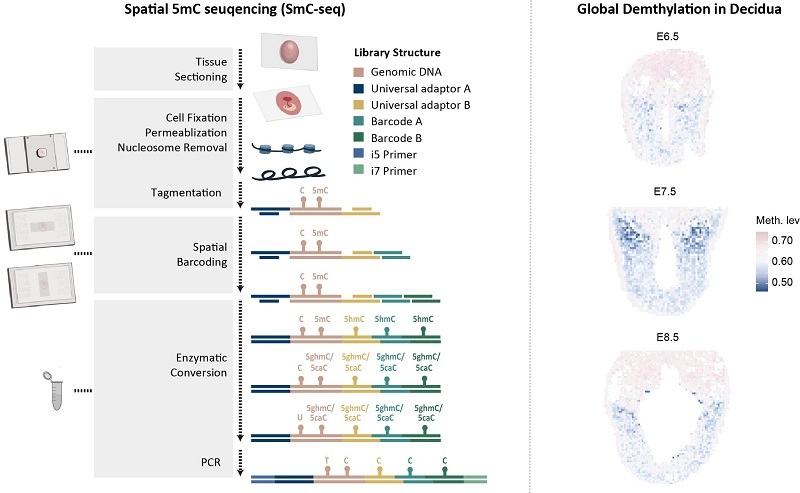
Figure: Principle of SmC-seq and global demethylation in maternal decidual cells
(Image by LIU Jiang's group)
Left panel: Schematic illustration of the SmC-seq (spatial methylation sequencing at single-cell scale) workflow. Key steps include tissue sectioning, sample pretreatment, spatial barcoding, and enzymatic methylation conversion.
Right panel: SmC-seq reveals that a subset of cells in the maternal uterine decidua exhibit global DNA demethylation.
Article link: https://www.nature.com/articles/s41592-026-03079-w
Contact: LIU Jiang
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
E-mail: liujiang@ibp.ac.cn
(Reported by Prof. LIU Jiang's group)

