Structural analysis for the post-fusion SARS-CoV spike glycoprotein
On July 17, 2020, a research paper led by Dr. ZHANG Xinzheng at Institute of Biophysics (IBP), entitled "Cryo-EM analysis of the post-fusion structure of the SARS-CoV spike glycoprotein", was published online in Nature Communications. In this work, the researchers used single-particle cryo-electron microscopy (Cryo-EM) to show the structural basis for the SARS-CoV spike (S) glycoprotein in the post-fusion state, which further reveals the fusion mechanism of coronaviruses and suggests potential targets for the development of vaccines and therapies that may suitable for treating a wide range of SARS-like coronaviruses.
Global emergencies caused by the highly pathogenic human-infecting coronaviruses have represented serious threats to human public health. However, effective vaccines and therapies are still lacking. Considering the potential risk of the spread of these or other unknown coronaviruses, as well as the relatively high mutation rates of RNA viral genomes, development of broad-spectrum antiviral treatments are required to prevent a wide range of coronaviruses infection. Coronavirus S glycoprotein, which mediates efficient transmission among species, is the major target of neutralizing antibodies and the key determinant for drug development. The S glycoprotein is a class I viral fusion protein that exists as a homotrimer on the virus surface, and its conserved S2 subunit is required for fusion of the membranes of the virus and its host cell. Most of the structural studies of S glycoproteins focused on the pre-fusion structures, and the structural information of the post-fusion S2 from these highly pathogenic human-infecting coronaviruses is still unknown.
In this work, Dr. ZHANG's group successfully solved the cryo-EM structure of the induced post-fusion SARS-CoV S2 machinery. The overall structure of the SARS-CoV S2 trimer adopts the shape of a baseball bat (Fig. 1), revealing large scale of structural rearrangements and conformational changes during membrane fusion. The HR1 motif turns over to form a continuous stem helix together with the CH motif pointing toward the target membrane, which makes the post-fusion SARS-CoV S2 fusion machinery extend ~80 angstrom longer that the pre-fusion S2 trimer. The researchers further found that the linker region upstream of the HR2 motif is released from the well-folded structure in the pre-fusion state, and extends along the axis of the stem helices, which is important for the formation of the central HR1-HR2 six-helix bundle in the post-fusion structure.
The coronavirus S glycoprotein contains a large number of N-linked glycans that cover the protein surface to limit its accessibility to neutralizing antibodies and evade attack by the host immune system. The structure presented here identified 24 out of 27 putative N-linked glycans (Fig. 1d), which reveal the distribution of the glycan shiled covering the surface of the post-fusion SARS-CoV S2 trimer. Since the amino acid sequences of the S2 subunits are highly conserved among different coronaviruses compared to that of the S1 subunits, therapeutics developed based on the S2 subunit may be more effective against a wide range of coronaviruses. Moreover, several S2-specific antibodies and inhibitors have been reported to show antiviral ability against a broader range of coronaviruses. The researchers mapped the published therapeutic targets within the S2 fusion machinery, and analyzed the potential targets that may suitable for broad-spectrum vaccines and drugs development against SARS-like coronaviruses.
Prof. ZHANG Xinzheng and Dr. CAO Duanfang are the co-corresponding authors. FAN Xiaoyi and Dr. CAO Duanfang are the co-first authors. KONG Lingfei participates in this work. The project was funded by National Natural Science Foundation of China and the Youth Innovation Promotion Association of the Chinese Academy of Sciences.
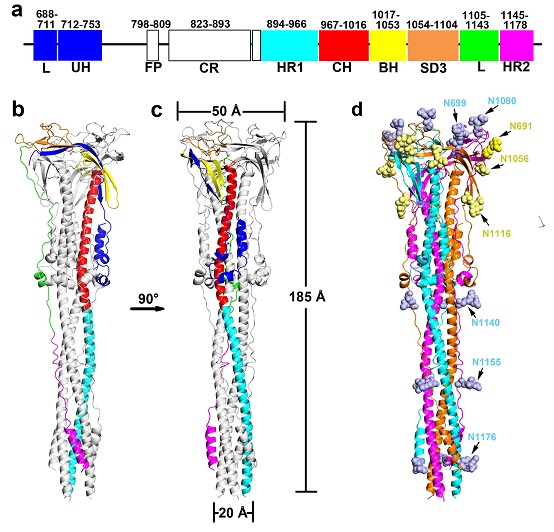
Figure 1. Overall structure and glycan shield of the post-fusion SARS-CoV S glycoprotein.
(Image by Dr. ZHANG Xinzheng’s group)
The web link for this paper is https://www.nature.com/articles/s41467-020-17371-6
Contact: ZHANG Xinzheng
Institute of Biophysices, Chinese Academy of Sciences
Beijing 100101, China
Email: xzzhang@ibp.ac.cn
(Reported by Dr. ZHANG Xinzheng’s group)

