Research reveals 3D landscape of Hepatitis B virus interactions with human chromatins
About 240 million people worldwide are chronically infected with Hepatitis B virus (HBV), a small enveloped DNA virus of 3.2 kb belonging to the Hepadnaviride family, which remains a serious global health problem. Uncontrolled chronic HBV infection can progress to life-threatening end-stage chronic liver diseases, such as cirrhosis and Hepatocellular carcinoma (HCC). The human nuclear HBV DNAs, including both covalently closed circular DNA (cccDNA) and integrated HBV DNA forms, are considered to be primary contributors to development and progression of HBV-associated liver diseases.
Prof. YANG Pengyuan at the Institute of Biophysics (IBP), Chinese Academy of Sciences (CAS) together with Prof. JI Xiong at School of Life Sciences in Beijing University emphasized the key roles of nuclear HBV DNAs in human cells and investigated how HBV DNAs communicate with human chromatin. This study was published in Cell Discovery online on December 29, 2020.
Researchers firstly employed a highly sensitive technology, 3C-HTGTS (3C-high-throughput genome-wide translocation sequencing), to globally identify HBV DNA-host DNA contacts in cellular models of HBV infection. Interestingly, HBV DNA does not randomly position in host genome, but instead preferentially establishes contacts with the host DNA at active chromatin regions. Importantly, the data from 3C-HTGTS revealed another kind of HBV DNA, integrated HBV DNA, could form chromatin loop with host genomic DNA at transcriptionally active region of human genome.
Based on the identification on the HBV DNA-host DNA contacts, researchers revealed H3K4me1-marked regions modified by KMT2C/D are also observed in the HBV cccDNA mini-chromosome and strongly influence HBV transcription. Furthermore, HBV infection influences host gene expression accompanied with HBV DNA-host DNA contacts. These findings provided a high throughput 3D landscape for the spatial organizations of cccDNA and integrated HBV DNA within the human genome, and a foundation to further understand the mechanisms of HBV modulation of liver disease development and progression in the future.
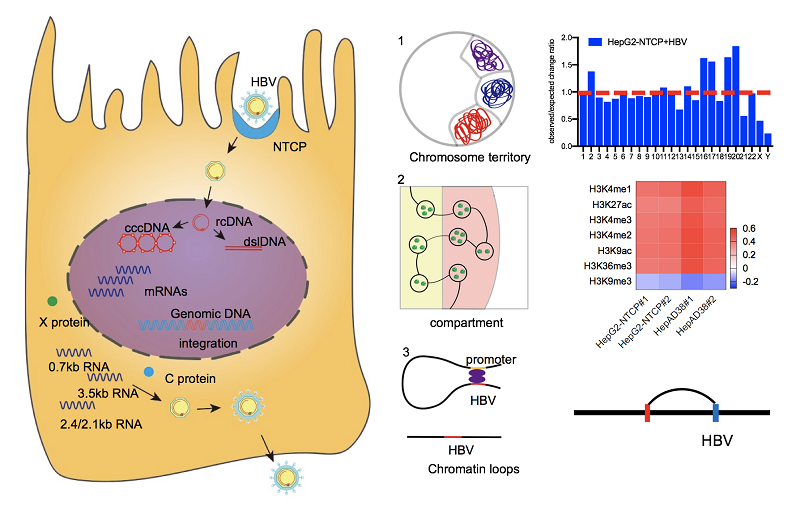
Figure 1. The model of the spatial organization of HBV DNAs in human nuclear.
(Image by Dr. YANG Pengyuan's group)
This project was supported by the National Natural Science Foundation of China, the Chinese Ministry of Science and Technology, and Chinese Academy of Sciences.
Article link: https://www.nature.com/articles/s41421-020-00218-1
Contact: YANG Pengyuan
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
Email: pyyang@ibp.ac.cn
(Reported by Dr. YANG Pengyuan's group)

