Dr. ZHU Bing's group published the protocol of Hammer-seq
On March 8, 2021, Nature Protocols published a protocol paper from ZHU Bing's lab, which is entitled " Simultaneously measuring the methylation of parent and daughter strands of replicated DNA at the single-molecule level by Hammer-seq ".
In the related previously published work (Ming et al., 2020, Cell Reserch), ZHU Bing's group developed Hammer-seq (Hairpin-assisted mapping of methylation of replicated DNA) to study the kinetics of DNA methylation maintenance and the underlying mechanisms. Hammer-seq combined 5-Ethynyl-2′-deoxyuridine (EdU) labeling, biotin conjugation by click reaction, streptavidin affinity purification and whole genome hairpin bisulfite-sequencing technologies.
For the convenience of other labs to use and advance Hammer-seq, we described the design, develop, steps, advantages and disadvantages of Hammer-seq in great detail in this Nature Protocols paper. Meanwhile, we suggested the potential applications of Hammer-seq in the future studies, which includes dissecting methylation maintenance kinetics, evaluating the accompanying de novo methylation events, and investigating the roles of specific proteins in regulating methylation maintenance. Notably, the simplified version of Hammer-seq can be an alternative and more accessible method to prepare the whole genome hairpin bisulfite-sequencing library.
This work is supported by the Ministry of Science and Technology of China, the National Natural Science Foundation of China, and the Chinese Academy of Sciences. Dr. ZHANG Zhuqiang is the corresponding author, Dr. MING Xuan is the first author, and Prof. ZHU Bing is also one of the authors of this paper. Dr. ZHANG Zhuqiang is also supported by the Youth Innovation Promotion Association of CAS.
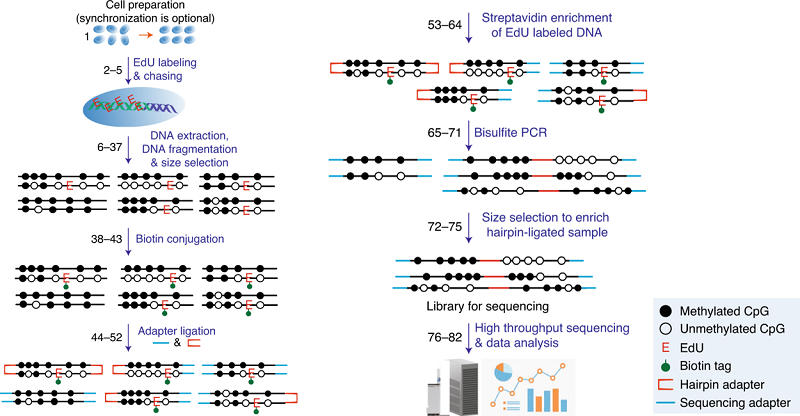
Schematic Overview of Hammer-seq.
The web link of this paper: https://www.nature.com/articles/s41596-020-00488-9
Contact: ZHANG Zhuqiang
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
Email: zhangzhuqiang@ibp.ac.cn
(Reported by Dr. ZHU Bing's group)

