Researchers Develop High Throughput Tool for Profiling Membrane Contact Sites in Cells
In a study published in Journal of Cell Biology, a deep learning-based high throughput quantification method for investigating organelle interactions was developed through the cooperation of Prof. XU Tao and Prof. HU Junjie at Institute of Biophysics and Dr. XIAO Li at Institute of Computing Technology, Chinese Academy of Sciences.
Membrane contact site (MCS)-mediated organelle interactions play essential roles in the cell. Quantitative analysis of MCSs has revealed vital clues for cellular responses under various physiological and pathological conditions. However, an efficient tool is lacking. The researchers developed “DeepContact”, a deep learning protocol for optimizing organelle segmentation and contact analysis based on label-free electron microscopy (EM). The system automates the processing of 2D EM slices, segmenting and visualizing MCSs in versatile width ranges, and quantifying normalized MCS parameters in large sample sizes, enabling cross-sample contact analysis.
DeepContact accommodates new morphologies of organelles and recognizes contacts in versatile width ranges, which enables statistical analysis of various types of MCSs in multiple systems. DeepContact profiled previously unidentified coordinative rearrangements of MCS types in cultured cells with combined nutritional conditioning. DeepContact also unveiled a subtle wave of ER-mitochondrial entanglement in Sertoli cells during the seminiferous epithelial cycle, indicating its potential in bridging MCS dynamics to physiological and pathological processes.
The code, models, and training data for DeepContact are open-source and can be incorporated into Amira for a friendly GUI interface for users with a computationally accessible setup.
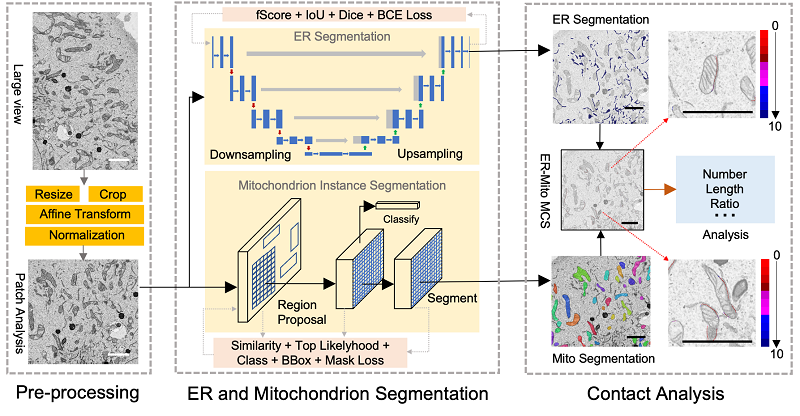
The DeepContact workflow
Article link:https://rupress.org/jcb/article/221/9/e202106190/213379/DeepContact-High-throughput-quantification-of?searchresult=1
Contact: LIU Liqing
Institute of Biophysics, Chinese Academy of Sciences
Beijing 100101, China
Email: liuliqing@ibp.ac.cn
(Reported by Dr. XU Tao's group)

